Glossary of genetics (M–Z)
This glossary of genetics is a list of definitions of terms and concepts commonly used in the study of genetics and related disciplines in biology, including molecular biology, cell biology, and evolutionary biology.[1] It is intended as introductory material for novices; for more specific and technical detail, see the article corresponding to each term. For related terms, see Glossary of evolutionary biology.
This glossary is split across two articles:
- Glossary of genetics (0–L) lists terms beginning with numbers and those beginning with the letters A through L.
- Glossary of genetics (M–Z) (this page) lists terms beginning with the letters M through Z.
M
- M phase
- See mitosis.
- macronucleus
- major groove
- map-based cloning
- See positional cloning.
- medical genetics
- The branch of medicine and medical science that involves the study, diagnosis, and management of hereditary disorders, and more broadly the application of knowledge about human genetics to medical care.
- meiosis
- A specialized type of cell division that occurs exclusively in sexually reproducing eukaryotes, during which DNA replication is followed by two consecutive rounds of division to ultimately produce four genetically unique haploid daughter cells, each with half the number of chromosomes as the original diploid parent cell. Meiosis only occurs in cells of the sex organs, and serves the purpose of generating haploid gametes such as sperm, eggs, or spores, which are later fused during fertilization. The two meiotic divisions, known as Meiosis I and Meiosis II, may also include various genetic recombination events between homologous chromosomes.
- meiotic spindle
- See spindle apparatus.
- melting
- The denaturation of a double-stranded nucleic acid into two single strands, especially in the context of the polymerase chain reaction.
- messenger RNA (mRNA)
- Any of a class of single-stranded RNA molecules which function as molecular messengers, carrying sequence information encoded in the DNA genome to the ribosomes where protein synthesis occurs. The primary products of transcription, mRNAs are synthesized by RNA polymerase, which builds a chain of ribonucleotides that complement the deoxyribonucleotides of a DNA template; in this way, the DNA sequence of a protein-coding gene is effectively preserved in the raw transcript, which is subsequently processed into a mature mRNA by a series of post-transcriptional modifications.
- metabolome
- metacentric
- (of a linear chromosome or chromosome fragment) Having a centromere positioned in the middle of the chromosome, resulting in chromatid arms of approximately equal length.[2]
- metaphase
- The stage of mitosis and meiosis that occurs after prometaphase and before anaphase, during which the centromeres of the replicated chromosomes align along the equator of the cell, with each kinetochore attached to the mitotic spindle.
- MicroArray and Gene Expression (MAGE)
- A group that "aims to provide a standard for the representation of DNA microarray gene expression data that would facilitate the exchange of microarray information between different data systems".[3]
- microchromosome
- A type of very small chromosome, generally less than 20,000 base pairs in size, present in the karyotypes of some organisms.
- microdeletion
- A chromosomal deletion that is too short to cause any apparent change in morphology under a light microscope, though it may still be detectable with other methods such as sequencing.
- micronucleus
- The smaller of the two types of nuclei that occur in pairs in the cells of some ciliated protozoa. Whereas the larger macronucleus is polyploid, the micronucleus is diploid and generally transcriptionally inactive.[4]
- microRNA (miRNA)
- A type of small, single-stranded, non-coding RNA molecule that functions in post-transcriptional regulation of gene expression, particularly RNA silencing, by base-pairing with complementary sequences in mRNA transcripts, which typically results in the cleavage or destabilization of the transcript or inhibits its translation by ribosomes.
- microsatellite
- A type of satellite DNA consisting of a relatively short sequence of tandem repeats, in which certain motifs (ranging in length from one to six or more bases) are repeated, typically 5–50 times. Microsatellites are widespread throughout most organisms' genomes and tend to have higher mutation rates than other regions. They are classified as variable number tandem repeat (VNTR) DNA, along with longer minisatellites.
- microspike
- See filopodium.
- microtomy
- microtrabecula
- A fine protein filament of the cytoskeleton. Multiple filaments form the microtrabecular network.[4]
- microtubule
- microtubule-organizing center
- microvillus
- mid body
- The centrally constricted region that forms across the central axis of a cell during cell division, constricted by the closing of the contractile ring until the daughter cells are finally separated.[4]
- middle lamella
- Minimum information about a microarray experiment (MIAME)
- A commercial standard developed by FGED and based on MAGE in order to facilitate the storage and sharing of gene expression data.[5][6]
- Minimal information about a high-throughput sequencing experiment (MINSEQE)
- A commercial standard developed by FGED for the storage and sharing of high-throughput sequencing data.[7]
- minisatellite
- A region of repetitive, non-coding genomic DNA in which certain DNA motifs (typically 10–60 bases in length) are tandemly repeated (typically 5–50 times). In the human genome, minisatellites occur at more than 1,000 loci, especially in centromeres and telomeres, and exhibit high mutation rates and high variability between individuals. Like the shorter microsatellites, they are classified as variable number tandem repeats (VNTRs) and are a type of satellite DNA.
- minor groove
- minus-strand
- See template strand.
- miRNA
- See microRNA.
- mismatch repair (MMR)
- mismatch
- An incorrect pairing of nucleobases on complementary strands of DNA or RNA, i.e. the presence in one strand of a duplex molecule of a base that is not complementary (by Watson–Crick pairing rules) to the base occupying the corresponding position in the other strand, which prevents normal hydrogen bonding between the bases; e.g. a guanine matched with a thymine.[8]
- missense mutation
- A type of point mutation which results in a codon that codes for a different amino acid than in the unmutated sequence. Compare nonsense mutation.
- mistranslation
- The insertion of an incorrect amino acid in a growing peptide chain during translation, i.e. the inclusion of any amino acid that is not the one specified by a particular codon in an mRNA transcript. Mistranslation may originate from a mischarged transfer RNA or from a malfunctioning ribosome.[8]
- mitochondrial DNA (mtDNA)
- mitochondrion
- mitosis
- In eukaryotic cells, the part of the cell cycle during which the division of the nucleus takes place and replicated chromosomes are separated into two distinct nuclei. Mitosis is generally preceded by the S phase of interphase, when the cell's DNA is replicated, and either occurs simultaneously with or is followed by cytokinesis, when the cytoplasm and plasma membrane are divided into two new daughter cells. Colloquially, the term "mitosis" is often used to refer to the entire process of cell division, not just that of the nucleus.
- mitotic index (MI)
- The proportion of cells within a sample which are undergoing mitosis at the time of observation, typically expressed as a percentage or as a value between 0 and 1. The number of cells dividing by mitosis at any given time can vary widely depending on organism, tissue, developmental stage, and culture media, among many other factors.[4]
- mitotic recombination
- mitotic segregation
- mitotic spindle
- See spindle apparatus.
- mixoploidy
- The presence of more than one different ploidy level, i.e. more than one number of sets of chromosomes, in different cells of the same cellular population.[8]
- mobile genetic element (MGE)
- Any genetic material that can move between different parts of a genome or be transferred from one species or replicon to another within a single generation. The many types of MGEs include transposable elements, bacterial plasmids, bacteriophage elements which integrate into host genomes by viral transduction, and self-splicing introns.
- mobilome
- The entire set of mobile genetic elements within a particular genome, cell, species, or other taxon, including all transposons, plasmids, prophages, and other self-splicing nucleic acid molecules.
- molecular cloning
- Any of various molecular biology methods designed to replicate a particular molecule, usually a DNA sequence, many times inside the cells of a natural host. Commonly, a recombinant DNA fragment containing a gene of interest is ligated into a plasmid vector, which is then transformed into competent bacterial cells. The bacteria, carrying the recombinant plasmid, are then allowed to proliferate naturally on a culture medium, where the plasmids are replicated along with the rest of the bacterial genome; any functioning gene of interest will be expressed by the bacterial cells, and thereby its gene products will also be cloned. Molecular cloning is a fundamental tool of genetic engineering, designed to study gene expression, to amplify a specific gene product, and to generate a selectable phenotype.
- molecular genetics
- A branch of genetics that employs methods and techniques of molecular biology to study the structure and function of genes and gene products at the molecular level. Contrast classical genetics.
- monad
- A haploid set of chromosomes as it exists inside the nucleus of an immature gametic cell such as an ootid or spermatid, i.e. a cell which has completed meiosis but is not yet a mature gamete.[8]
- monocentric
- (of a linear chromosome or chromosome fragment) Having only one centromere. Contrast dicentric and holocentric.
- monoclonal antibody (mAb)
- monoploid
- monosomy
- The abnormal and frequently pathological presence of only one chromosome of a normal diploid pair. It is a type of aneuploidy.
- Morpholino
- A synthetic polymeric molecule connecting short sequences of nucleobases into artificial antisense oligomers, which are used in genetic engineering to knockdown gene expression by pairing with complementary sequences in naturally occurring RNA or DNA molecules, especially mRNA transcripts, thereby blocking access by other biomolecules such as ribosomes and proteins. Morpholino oligomers are not themselves translated, and neither they nor their hybrid duplexes with RNA are attacked by nucleases; also, unlike the negatively charged phosphates of normal nucleic acids, the synthetic backbones of Morpholinos are electrically neutral, making them less likely to interact non-selectively with a host cell's charged proteins. These properties make them useful and reliable tools for artificially generating mutant phenotypes in living cells.[8]
- mosaicism
- The presence of two or more populations of cells with different genotypes in an individual organism which has developed from a single fertilized egg. A mosaic organism can result from many kinds of genetic phenomena, including nondisjunction of chromosomes, endoreduplication, or mutations in individual stem cell lineages during the early development of the embryo. Mosaicism is similar to but distinct from chimerism.
- motif
- Any distinctive or recurring sequence of nucleotides in a nucleic acid or of amino acids in a protein that is or is conjectured to be biologically significant, especially one that is reliably recognized by other biomolecules or which has a three-dimensional structure that permits unique or characteristic chemical interactions such as DNA binding.[8] In nucleic acids, motifs are often short (three to ten nucleotides in length), highly conserved sequences which act as recognition sites for DNA-binding enzymes or RNAs involved in the regulation of gene expression.
- motor protein
- mRNA
- See messenger RNA.
- mtDNA
- See mitochondrial DNA.
- multinucleate
- (of a cell) Having more than one nucleus within a single cell; i.e. having multiple nuclei occupying the same cytoplasm.
- multiomics
- multiple cloning site (MCS)
- mutagen
- Any physical or chemical agent that changes the genetic material (usually DNA) of an organism and thereby increases the frequency of mutations above natural background levels.
- mutagenesis
- 1. The process by which the genetic information of an organism is changed, resulting in a mutation. Mutagenesis may occur spontaneously or as a result of exposure to a mutagen.
- 2. In molecular biology, any laboratory technique by which one or more genetic mutations are deliberately engineered in order to produce a mutant gene, regulatory element, gene product, or genetically modified organism so that the functions of a genetic locus, process, or product can be studied in detail.
- mutant
- An organism, gene product, or phenotypic trait resulting from a mutation, of a type that would not be observed naturally in wild-type specimens.
- mutation
- Any permanent change in the nucleotide sequence of a strand of DNA or RNA, or in the amino acid sequence of a peptide. Mutations play a role in both normal and abnormal biological processes; their natural occurrence is integral to the process of evolution. They can result from errors in replication, chemical damage, exposure to high-energy radiation, or manipulations by mobile genetic elements. Repair mechanisms have evolved in many organisms to correct them. By understanding the effect that a mutation has on phenotype, it is possible to establish the function of the gene or sequence in which it occurs.
- mutator gene
- Any mutant gene or sequence that increases the spontaneous mutation rate of one or more other genes or sequences. Mutators are often transposable elements, or may be mutant housekeeping genes such as those that encode helicases or proteins involved in proofreading.[8]
- mutein
- A mutant protein, i.e. a protein whose amino acid sequence differs from that of the normal because of a mutation.
- muton
- The smallest unit of a DNA molecule in which a physical or chemical change can result in a mutation (conventionally a single nucleotide).[8]

The structure of a typical mature protein-coding messenger RNA or mRNA, drawn approximately to scale. The coding sequence (green) is bounded by untranslated regions at both the 5'-end (yellow) and the 3'-end (pink). Prior to export from the nucleus, a 5' cap (red) and a 3' poly(A) tail (black) are added to help stabilize the mRNA and prevent its degradation by ribonucleases.
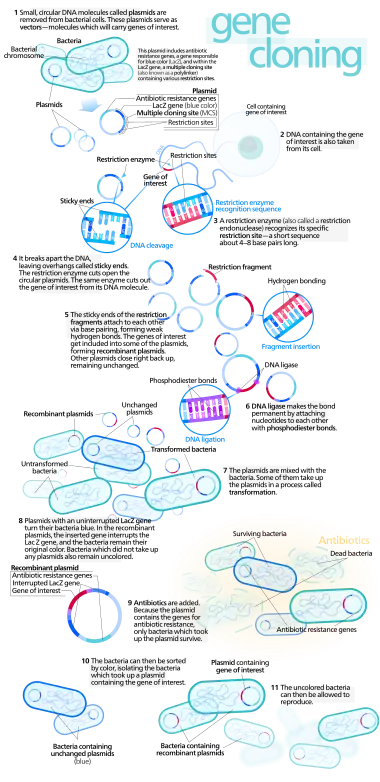
An outline of how molecular cloning works
N
- n orientation
- One of two possible orientations by which a linear DNA fragment can be inserted into a vector, specifically the one in which the gene maps of both fragment and vector have the same orientation.[8] Contrast u orientation.
- N-terminus
- The end of a linear chain of amino acids (i.e. a peptide) that is terminated by the free amine group (–NH
2) of the first amino acid added to the chain during translation. This amino acid is said to be N-terminal. By convention, sequences, domains, active sites, or any other structure positioned nearer to the N-terminus of the polypeptide or the folded protein it forms relative to others are described as upstream. Contrast C-terminus. - NAD
- See nicotinamide-adenine dinucleotide.
- NADP
- See nicotinamide-adenine dinucleotide phosphate.
- nascent
- In the process of being synthesized; incomplete; not yet fully processed or mature. The term is commonly used to describe strands of DNA or RNA which are actively undergoing synthesis during replication or transcription, respectively, or sometimes a complete, fully transcribed RNA molecule before any alterations have been made (e.g. polyadenylation or RNA editing), or a peptide chain actively undergoing translation by a ribosome.[8]
- ncDNA
- See non-coding DNA.
- ncRNA
- See non-coding RNA.
- negative (-) sense strand
- See template strand.
- negative control
- A type of gene regulation in which a repressor binds to an operator upstream from the coding region and thereby prevents transcription by RNA polymerase. An inducer is necessary to switch on transcription in positive control.[9]
- negative supercoiling
- nick
- nick translation
- nicking enzyme
- nicotinamide adenine dinucleotide (NAD)
- nicotinamide adenine dinucleotide phosphate (NADP+, NADP)
- nitrogenous base
- Any organic compound containing a nitrogen atom that has the chemical properties of a base. A set of five particular nitrogenous bases – adenine (A), guanine (G), cytosine (C), thymine (T), and uracil (U) – are especially relevant to biology because they are components of nucleotides, which in turn are the primary monomers that make up nucleic acids.
- non-canonical amino acid (ncAA)
- Any amino acid, natural or artificial, that is not one of the 20 or 21 proteinogenic amino acids encoded by the standard genetic code. There are hundreds of such amino acids, many of which have biological functions and are specified by alternative codes or incorporated into proteins accidentally by errors in translation. Many of the best known naturally occurring ncAAs occur as intermediates in the metabolic pathways leading to the standard amino acids, while others have been made synthetically in the laboratory.[10]
- non-coding DNA (ncDNA)
- Any segment of DNA that does not encode a sequence that may ultimately be transcribed and translated into a protein. In most organisms, only a small fraction of the genome consists of protein-coding DNA, though the proportion varies greatly between species. Some non-coding DNA may still be transcribed into functional non-coding RNA (as with transfer RNAs) or may serve important developmental or regulatory purposes; other regions (as with so-called "junk DNA") appear to have no known biological function.
- non-coding RNA (ncRNA)
- Any molecule of RNA that is not ultimately translated into a protein. The DNA sequence from which a functional non-coding RNA is transcribed is often referred to as an "RNA gene". Numerous types of non-coding RNAs essential to normal genome function are produced constitutively, including transfer RNA (tRNA), ribosomal RNA (rRNA), microRNA (miRNA), and small interfering RNA (siRNA); other non-coding RNAs (sometimes described as "junk RNA") have no known function and are likely the product of spurious transcription.
- non-homologous end joining (NHEJ)
- non-transcribed spacer (NTS)
- See spacer.
- noncoding strand
- See template strand.
- nondisjunction
- The failure of homologous chromosomes or sister chromatids to segregate properly during cell division. Nondisjunction results in daughter cells that are aneuploid, containing abnormal numbers of one or more specific chromosomes. It may be caused by any of a variety of factors.
- nonrepetitive sequence
- Broadly, any nucleotide sequence or region of a genome that does not contain repeated sequences, or in which repeats do not comprise a majority; or any segment of DNA exhibiting the reassociation kinetics expected of a unique sequence.[8]
- nonsense mutation
- A type of point mutation which results in a premature stop codon in the transcribed mRNA sequence, thereby causing the premature termination of translation, which results in a truncated, incomplete, and often non-functional protein.
- nonsense suppressor
- nonsynonymous mutation
- A type of mutation in which the substitution of one nucleotide base for another results, after transcription and translation, in an amino acid sequence that is different from that produced by the original unmutated gene. Because nonsynonymous mutations always result in a biological change in the organism, they are often subject to strong selection pressure. Contrast synonymous mutation.
- northern blotting
- A blotting method in molecular biology used to detect RNA in a sample. Compare Southern blotting, western blotting, and eastern blotting.
- nRNA
- See nuclear RNA.
- nuclear cage
- nuclear envelope
- A sub-cellular barrier consisting of two concentric lipid bilayer membranes that surrounds the nucleus in eukaryotic cells. The nuclear envelope is sometimes simply called the "nuclear membrane", though the structure is actually composed of two distinct membranes, an inner membrane and an outer membrane.
- nuclear pore
- nuclear membrane
- A sub-cellular barrier consisting of two lipid bilayer membranes that surrounds the nucleus in eukaryotic cells.
- nuclear RNA (nRNA)
- Any RNA molecule located within a cell's nucleus, whether associated with chromosomes or existing freely in the nucleoplasm, including small nuclear RNA (snRNA), enhancer RNA (eRNA), and all newly transcribed immature RNAs, coding or non-coding, prior to their export to the cytosol (hnRNA).
- nuclear transfer
- nuclease
- Any of a class of enzymes capable of cleaving phosphodiester bonds connecting adjacent nucleotides in a nucleic acid molecule (the opposite of a ligase). Nucleases may nick one or cut both strands of a duplex molecule; they may cleave randomly or at specific recognition sequences. They are ubiquitous and imperative for normal cellular function, and are also widely employed in molecular biology techniques.
- nucleic acid
- A long, polymeric macromolecule made up of smaller monomers called nucleotides which are chemically linked to one another in a chain. Two specific types of nucleic acid, DNA and RNA, are common to all living organisms, serving to encode the genetic information governing the construction, development, and ordinary processes of all biological systems. This information, contained within the order or sequence of the nucleotides, is translated into proteins, which direct all of the chemical reactions necessary for life.
- nucleic acid sequence
- The precise order of consecutively linked nucleotides in a nucleic acid molecule such as DNA or RNA. Long sequences of nucleotides are the principal means by which biological systems store genetic information, and therefore the accurate replication, transcription, and translation of such sequences is of the utmost importance, lest the information be lost or corrupted. Nucleic acid sequences may be equivalently referred to as sequences of nitrogenous bases, nucleobases, nucleotides, or base pairs, and they correspond directly to sequences of codons and amino acids.
- nucleobase
- Any of the five primary or canonical nitrogenous bases – adenine (A), guanine (G), cytosine (C), thymine (T), and uracil (U) – that form nucleosides and nucleotides, the latter of which are the fundamental building blocks of nucleic acids. The ability of these bases to form base pairs via hydrogen bonding, as well as their flat, compact three-dimensional profiles, allows them to "stack" one upon another and leads directly to the long-chain structures of DNA and RNA. When writing sequences in shorthand notation, the letter N is often used to represent a nucleotide containing a generic or unidentified nucleobase.
- nucleoid
- An irregularly shaped region within a prokaryotic cell which contains most or all of the cell's genetic material, but is not enclosed by a nuclear membrane as in eukaryotes.
- nucleolonema
- nucleolus
- An organelle within the nucleus of eukaryotic cells which is composed of proteins, DNA, and RNA and serves as the site of ribosome synthesis.
- nucleoplasm
- nucleoprotein
- Any protein that is chemically bonded to or conjugated with a nucleic acid molecule. Examples include ribosomes, nucleosomes, and many enzymes.
- nucleosidase
- Any of a class of enzymes which catalyze the decomposition of nucleosides into their component nitrogenous bases and pentose sugars.[8]
- nucleoside
- An organic molecule composed of a nitrogenous base bonded to a five-carbon sugar (either ribose or deoxyribose). A nucleotide additionally includes one or more phosphate groups.
- nucleosome
- The basic structural subunit of chromatin used in packaging nuclear DNA such as chromosomes, consisting of a core particle of eight histone proteins around which double-stranded DNA is wrapped in a manner akin to thread wound around a spool. The technical definition of a nucleosome includes a segment of DNA about 146 base pairs in length which makes 1.67 left-handed turns as it coils around the histone core, as well as a stretch of linker DNA (generally 38–80 bp) connecting it to an adjacent core particle, though the term is often used to refer to the core particle alone. Long series of nucleosomes are further condensed by association with histone H1 into higher-order structures such as 30-nm fibers and ultimately supercoiled chromatids. Because the histone–DNA interaction limits access to the DNA molecule by other proteins and RNAs, the precise positioning of nucleosomes along the DNA sequence plays a fundamental role in controlling whether or not genes are transcribed and expressed, and hence mechanisms for moving and ejecting nucleosomes have evolved as a means of regulating the expression of particular loci.
- nucleosome-depleted region (NDR)
- A region of a genome or chromosome in which long segments of DNA are bound by few or no nucleosomes and thus exposed to manipulation by other proteins and molecules, especially implying that the region is transcriptionally active.
- nucleotide
- An organic molecule that serves as the monomer or subunit of nucleic acid polymers, including RNA and DNA. Each nucleotide is composed of three connected functional groups: a nitrogenous base, a five-carbon sugar (either ribose or deoxyribose), and a single phosphate group. Though technically distinct, the term "nucleotide" is often used interchangeably with nitrogenous base, nucleobase, and base pair when referring to the sequences that make up nucleic acids. Compare nucleoside.
- nucleotide sequence
- See nucleic acid sequence.
- nucleus
- A large spherical or lobular organelle surrounded by a nuclear membrane which functions as the main storage compartment for the genetic material of eukaryotic cells, including the DNA comprising chromosomes, as well as the site of RNA synthesis during transcription. The vast majority of eukaryotic cells have a single nucleus, though some cells may have more than one nucleus, either temporarily or permanently, and in some organisms there exist certain cell types (e.g. mammalian erythrocytes) which lose their nuclei upon reaching maturity, effectively becoming anucleate. The nucleus is one of the defining features of eukaryotes; the cells of prokaryotes such as bacteria lack nuclei entirely.[4]
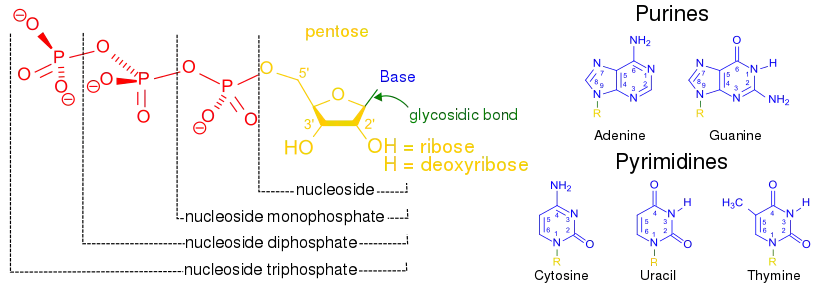
The nucleobases (blue) are the five specific nitrogenous bases canonically used in DNA and RNA. A nucleobase bonded to a pentose sugar (either ribose or deoxyribose; yellow) is known as a nucleoside (yellow + blue). A nucleoside bonded to a single phosphate group (red) is known as a nucleoside monophosphate (NMP) or a nucleotide (red + yellow + blue). When not incorporated into a nucleic acid chain, free nucleosides can bind multiple phosphate groups: two phosphates yields a nucleoside diphosphate (NDP), and three yields a nucleoside triphosphate (NTP).
O
- occluding junction
- ochre
- One of three stop codons used in the standard genetic code; in RNA, it is specified by the nucleotide triplet UAA. The other two stop codons are named amber and opal.
- Okazaki fragments
- Short sequences of nucleotides which are synthesized discontinuously by DNA polymerase and later linked together by DNA ligase to create the lagging strand during DNA replication. Okazaki fragments are the consequence of the unidirectionality of DNA polymerase, which only works in the 5' to 3' direction.
- oligogene
- oligomer
- Any polymeric molecule consisting of a relatively short series of connected monomers or subunits; e.g. an oligonucleotide is a short series of nucleotides.
- oligonucleotide
- A relatively short chain of nucleic acid residues. In the laboratory, oligonucleotides are commonly used as primers or hybridization probes to detect the presence of larger mRNA molecules or assembled into two-dimensional microarrays for high-throughput sequencing analysis.
- oligosaccharide
- omnipotent suppressor
- oncogene
- A gene that has the potential to cause cancer. In tumor cells, such genes are often mutated and/or expressed at abnormally high levels.
- one gene–one polypeptide
- The hypothesis that there exists a large class of genes in which each particular gene directs the synthesis of one particular polypeptide or protein.[8] Historically it was thought that all genes and proteins might follow this rule by definition, but it is now known that many or most proteins are composites of different polypeptides and therefore the product of multiple genes, and also that some genes do not encode polypeptides at all but instead produce non-coding RNAs, which are never translated.
- opal
- One of three stop codons used in the standard genetic code; in RNA, it is specified by the nucleotide triplet UGA. The other two stop codons are named amber and ochre.
- open chromatin
- See euchromatin.
- open reading frame (ORF)
- The part of a reading frame that has the ability to be translated from DNA or RNA into protein; any continuous stretch of codons that contains a start codon and a stop codon.
- operator
- A regulatory sequence within an operon, typically located between the promoter sequence and the structural genes of the operon, to which an uninhibited repressor protein can bind, thereby physically obstructing RNA polymerase from initiating the transcription of adjacent cistrons.[11]
- operon
- A functional unit of gene expression consisting of a cluster of adjacent structural genes which are collectively under the control of a single promoter, along with one or more adjacent regulatory sequences such as operators which affect transcription of the structural genes. The set of genes is transcribed together, usually resulting in a single polycistronic messenger RNA molecule, which may then be translated together or undergo splicing to create multiple mRNAs which are translated independently; the result is that the genes contained in the operon are either expressed together or not at all. Regulatory proteins, including repressors, corepressors, and activators, usually bind specifically to the regulatory sequences of a given operon; by some definitions, the genes that code for these regulatory proteins are also considered part of the operon.
- operon network
- organelle
- origin of replication (ORI)
- A particular location within a DNA molecule at which DNA replication is initiated. Origins are usually defined by the presence of a replicator sequence or by specific chromatin patterns.
- outron
- A sequence near the 5'-end of a primary mRNA transcript that is removed by a special form of splicing during post-transcriptional processing. Outrons are located entirely outside of the transcript's coding sequences, unlike introns.
- overexpression
- An abnormally high level of gene expression which results in an excessive number of copies of one or more gene products. Overexpression produces a pronounced gene-related phenotype.[12][13]
- oxidative phosphorylation
P
- p53
- pachynema
- In meiosis, the third of five substages of prophase I, following zygonema and preceding diplonema. During pachynema, the synaptonemal complex facilitates crossing over between the synapsed homologous chromosomes, and the centrosomes begin to move apart from each other.[8]
- palindromic sequence
- A nucleic acid sequence of a double-stranded DNA or RNA molecule in which the unidirectional sequence (e.g. 5' to 3') of nucleotides on one strand matches the sequence in the same direction (e.g. 5' to 3') on the complementary strand. In other words, a nucleotide sequence is said to be palindromic if it is equal to its own reverse complement. Palindromic motifs are common in most genomes and are capable of forming hairpins.
- passenger
- A DNA fragment of interest designed to be spliced into a 'vehicle' such as a plasmid vector and then cloned.[8]
- PCR
- See polymerase chain reaction.
- PCR product
- See amplicon.
- peptide
- A short chain of amino acid monomers linked by covalent peptide bonds. Peptides are the fundamental building blocks of longer polypeptide chains and hence of proteins.
- peptide bond
- A covalent chemical bond between the carboxyl group of one amino acid and the amino group of an adjacent amino acid in a peptide chain, formed by a dehydration reaction catalyzed by peptidyl transferase, an enzyme within the ribosome, during translation.
- perinuclear space
- The space between the inner and outer membranes of the nuclear envelope.
- peroxisome
- pervasive transcription
- phagocyte
- A type of cell which functions as part of the immune system by engulfing and ingesting harmful foreign molecules, bacteria, and dead or dying cells, in a process known as phagocytosis.
- phagocytosis
- phagosome
- pharmacogenomics
- The study of the role played by the genome in the body's response to pharmaceutical drugs, combining the fields of pharmacology and genomics.
- phenome
- The complete set of phenotypes that are or can be expressed by a genome, cell, tissue, organism, or species; the sum of all of its manifest chemical, morphological, and behavioral characteristics or traits.
- phenomic lag
- A delay in the phenotypic expression of a genetic mutation owing to the time required for the manifestation of changes in the affected biochemical pathways.[9]
- phenotype
- The composite of the observable morphological, physiological, and behavioral traits of an organism that result from the expression of the organism's genotype as well as the influence of environmental factors and the interactions between the two.
- phosphatase
- Any of a class of enzymes that catalyze the hydrolytic cleavage of a phosphoric acid monoester into a phosphate ion and an alcohol, e.g. the removal of a phosphate group from a nucleotide via the breaking of the ester bond connecting the phosphate to a ribose or deoxyribose sugar or to another phosphate, a process termed dephosphorylation. The opposite process is performed by kinases.
- phosphate backbone
- The linear chain of alternating phosphate and sugar compounds that results from the linking of consecutive nucleotides in the same strand of a nucleic acid molecule, and which serves as the structural framework of the nucleic acid. Each individual strand is held together by a repeating series of phosphodiester bonds connecting each phosphate group to the ribose or deoxyribose sugars of two adjacent nucleotides. These bonds are created by ligases and broken by nucleases.
- phosphodiester bond
- A pair of ester bonds linking a phosphate molecule with the two pentose rings of consecutive nucleosides on the same strand of a nucleic acid. Each phosphate forms a covalent bond with the 3' carbon of one pentose and the 5' carbon of the adjacent pentose; the repeated series of such bonds that holds together a long chain of nucleotides in DNA and RNA is known as the phosphate or phosphodiester backbone.
- phospholipid bilayer
- piRNA
- See Piwi-interacting RNA.
- Piwi-interacting RNA (piRNA)
- plasma membrane
- plasmid
- Any small DNA molecule that is physically separated from the larger body of chromosomal DNA and can replicate independently. Plasmids are most commonly found as small, circular, double-stranded DNA molecules in prokaryotes such as bacteria, though they are also sometimes present in archaea and eukaryotes.
- plastid
- ploidy
- The number of complete sets of chromosomes in a cell, and hence the number of possible alleles present within the cell at any given autosomal locus.
- pluripotency
- plus-strand
- See coding strand.
- point mutation
- A mutation by which a single nucleotide base is changed, inserted, or deleted from a sequence of DNA or RNA.
- poly(A) tail
- polyadenylation
- The addition of a series of multiple adenosine ribonucleotides, known as a poly(A) tail, to the 3'-end of a primary RNA transcript, typically a messenger RNA. A class of post-transcriptional modification, polyadenylation serves different purposes in different cell types and organisms. In eukaryotes, the addition of a poly(A) tail is an important step in the processing of a raw transcript into a mature mRNA, ready for export to the cytoplasm where translation occurs; in many bacteria, polyadenylation has the opposite function, instead promoting the RNA's degradation.
- polylinker
- See multiple cloning site.
- polymerase
- Any of a class of enzymes which catalyze the synthesis of polymeric molecules, especially nucleic acid polymers, typically by encouraging the pairing of free nucleotides to those of an existing complementary template. DNA polymerases and RNA polymerases are essential for DNA replication and transcription, respectively.
- polymerase chain reaction (PCR)
- Any of a wide variety of molecular biology methods involving the rapid production of millions or billions of copies of a specific DNA sequence, allowing scientists to selectively amplify fragments of a very small sample to a quantity large enough to study in detail. In its simplest form, PCR generally involves the incubation of a target DNA sample of known or unknown sequence with a reaction mixture consisting of oligonucleotide primers, a heat-stable DNA polymerase, and free deoxyribonucleotide triphosphates (dNTPs), all of which are supplied in excess. This mixture is then alternately heated and cooled to pre-determined temperatures for pre-determined lengths of time according to a specified pattern which is repeated for many cycles, typically in a thermal cycler which automatically controls the required temperature variations. In each cycle, the most basic of which includes a denaturation phase, annealing phase, and elongation phase, the copies synthesized in the previous cycle are used as templates for synthesis in the next cycle, causing a chain reaction that results in the exponential growth of the total number of copies in the reaction mixture. Amplification by PCR has become a standard technique in virtually all molecular biology laboratories.
- polypeptide
- A long, continuous, and unbranched polymeric chain of amino acid monomers linked by covalent peptide bonds, typically longer than a peptide. Proteins generally consist of one or more polypeptides arranged in a biologically functional way.
- polyploid
- (of a cell or organism) Having more than two homologous copies of each chromosome; i.e. any ploidy level that is greater than diploid. Polyploidy may occur as a normal condition of chromosomes in certain cells or even entire organisms, or it may result from errors in cell division or mutations causing the duplication of the entire chromosome set.
- polysaccharide
- polysome
- A complex of a messenger RNA molecule and two or more ribosomes which act to translate the mRNA transcript into a polypeptide.
- polysomy
- The condition of a cell or organism having at least one more copy of a particular chromosome than is normal for its ploidy level, e.g. a diploid organism with three copies of a given chromosome is said to show trisomy. Every polysomy is a type of aneuploidy.
- position effect
- Any effect on the expression or functionality of a gene or sequence that is a consequence of its location or position within a chromosome or other DNA molecule. A sequence's precise location relative to other sequences and structures tends to strongly influence its activity and other properties, because different loci on the same molecule can have substantially different genetic backgrounds and physical/chemical environments, which may also change over time. For example, the transcription of a gene located very close to a nucleosome, centromere, or telomere is often repressed or entirely prevented because the proteins that make up these structures block access to the DNA by transcription factors, while the same gene is transcribed at a much higher rate when located in euchromatin. Proximity to promoters, enhancers, and other regulatory elements, as well as to regions of frequent transposition by mobile elements, can also directly affect expression; being located near the end of a chromosomal arm or to common crossover points may affect when replication occurs and the likelihood of recombination. Position effects are a major focus of research in the field of epigenetic inheritance.
- positional cloning
- A strategy for identifying and cloning a candidate gene based on knowledge of its locus or position alone and with little or no information about its products or function, in contrast to functional cloning. This method usually begins by comparing the genomes of individuals expressing a phenotype of unknown provenance (often a hereditary disease) and identifying genetic markers shared between them. Regions defined by markers flanking one or more genes of interest are cloned, and the genes located between the markers can then be identified by any of a variety of means, e.g. by sequencing the region and looking for open reading frames, by comparing the sequence and expression patterns of the region in mutant and wild-type individuals, or by testing the ability of the putative gene to rescue a mutant phenotype.[8]
- positive (+) sense strand
- See coding strand.
- post-transcriptional modification
- post-translational modification
- primary transcript
- The unprocessed, single-stranded RNA molecule produced by the transcription of a DNA sequence as it exists before post-transcriptional modifications such as alternative splicing convert it into a mature RNA product such as an mRNA, tRNA, or rRNA. A precursor mRNA or pre-mRNA, for example, is a type of primary transcript that becomes a mature mRNA ready for translation after processing.
- primer
- A short, single-stranded oligonucleotide, typically 5–100 bases in length, which "primes" or initiates nucleic acid synthesis by hybridizing to a complementary sequence on a template strand and thereby provides an existing 3'-end from which a polymerase can extend the new strand. Natural systems exclusively use RNA primers to initiate DNA replication and transcription, whereas the in vitro syntheses performed in many laboratory techniques such as PCR often use DNA primers. In modern laboratories, primers are carefully designed, often in "forward" and "reverse" pairs, to complement specific and unique sequences within genomic DNA, with consideration given to their melting and annealing temperatures, and then purchased from commercial suppliers which create oligonucleotides on demand by de novo synthesis.
- primer dimer (PD)
- priming
- The initiation of nucleic acid synthesis by the hybridization or annealing of one or more primers to a complementary sequence within a template strand.
- probe
- Any reagent used to make a single measurement in a biochemical assay such as a gene expression experiment. Compare reporter.
- probe-set
- A collection of two or more probes designed to measure a single molecular species, such as a collection of oligonucleotides designed to hybridize to various parts of the mRNA transcripts generated from a single gene.
- process molecular gene concept
- prometaphase
- The second stage of cell division in mitosis, following prophase and preceding metaphase, during which the nuclear membrane disintegrates, the chromosomes inside form kinetochores around their centromeres, microtubules emerging from the poles of the mitotic spindle reach the nuclear space and attach to the kinetochores, and motor proteins associated with the microtubules begin to push the chromosomes toward the center of the cell.
- promoter
- A sequence or region of DNA, usually 100–1,000 base pairs long, to which transcription factors bind in order to recruit RNA polymerase to the sequence and initiate the transcription of one or more genes. Promoters are located upstream of the genes they transcribe, near the transcription start site.
- promotion
- See upregulation.
- prophase
- The first stage of cell division in both mitosis and meiosis, occurring after interphase and before prometaphase, during which the DNA of the chromosomes is condensed into chromatin, the nucleolus disintegrates, centrosomes move to opposite ends of the cell, and the mitotic spindle forms.
- protein
- A polymeric macromolecule composed of one or more long chains of amino acids linked by peptide bonds. Proteins are the three-dimensional structures created when these chains fold into specific higher-order arrangements following translation, and it is this folded structure which determines a protein's chemical activity and hence its biological function. Ubiquitous and fundamental in all living organisms, proteins are the primary means by which the activities of life are performed, participating in the vast majority of the biochemical reactions that occur inside and outside of cells. They are often classified according to the type(s) of reaction(s) they facilitate or catalyze, by the chemical substrate(s) they act upon, or by their functional role in cellular activity; e.g. as structural proteins, motor proteins, enzymes, transcription factors, or links within biochemical pathways.
- proteinogenic amino acid
- Any of the 20 canonical amino acids which are encoded by the standard genetic code and incorporated into peptides and ultimately proteins during translation. The term may also be inclusive of an additional two amino acids encoded by non-standard codes which can be incorporated by special translation mechanisms.
- proteome
- The entire set of proteins that is or can be expressed by a particular genome, cell, tissue, or species at a particular time (such as during a single lifespan or during a specific developmental stage) or under particular conditions (such as when compromised by a certain disease).
- proton motive force
- purine
- A double-ringed heterocyclic organic compound which, along with pyrimidine, is one of two molecules from which all nitrogenous bases (including the nucleobases used in DNA and RNA) are derived. Adenine (A) and guanine (G) are classified as purines. The letter R is sometimes used to indicate a generic purine; e.g. in a nucleotide sequence read, R may be used to indicate that either purine nucleobase, A or G, can be substituted at the indicated position.
- putative gene
- A specific nucleotide sequence suspected to be a functional gene based on the identification of its open reading frame. The gene is said to be "putative" in the sense that no function has yet been described for its products.
- pyrimidine
- A single-ringed heterocyclic organic compound which, along with purine, is one of two molecules from which all nitrogenous bases (including the nucleobases used in DNA and RNA) are derived. Cytosine (C), thymine (T), and uracil (U) are classified as pyrimidines. The letter Y is sometimes used to indicate a generic pyrimidine; e.g. in a nucleotide sequence read, Y may be used to indicate that either pyrimidine nucleobase – C, T, or U – can be substituted at the indicated position.
- pyrimidine dimer
- A type of molecular lesion caused by photochemical damage to DNA or RNA, whereby exposure to ultraviolet (UV) radiation induces the formation of covalent bonds between pyrimidine bases occupying adjacent positions in the same polynucleotide strand, which in turn may cause local conformational changes in secondary structure and prevent base pairing with the opposite strand. In DNA, the dimerization reaction occurs between neighboring thymine and cytosine residues (T−T, C−C, or T−C); it can also occur between cytosine and uracil residues in double-stranded RNA. Pyrimidine dimers are usually quickly corrected by nucleotide excision repair, but uncorrected lesions can inhibit or arrest polymerase activity during transcription or replication.
- pyruvic acid
Q
- quantitative PCR (qPCR)
- quiescent culture
- A cell culture in which there is little or no active cell growth or replication but in which the cells nonetheless continue to survive, as observed with some confluent cultures.[4]
R
- rDNA
- 1. An abbreviation of recombinant DNA.
- 2. An abbreviation of ribosomal DNA.
- reading frame
- A way of dividing the nucleotide sequence in a DNA or RNA molecule into a set of consecutive, non-overlapping triplets, which is "read" by proteins during transcription and replication. In coding DNA, each triplet is referred to as a codon that corresponds to a particular amino acid during translation. In general, only one reading frame (the so-called open reading frame) in a given section of a nucleic acid can be used to make functional proteins, but there are exceptions in a few organisms. A frameshift mutation results in a shift in the normal reading frame and affects all downstream codons.
- real-time PCR (rtPCR)
- See quantitative PCR.
- reassociation kinetics
- The measurement and manipulation of the rate of reannealing of complementary strands of DNA, generally by heating and denaturing a double-stranded molecule into single strands and then observing their rehybridization at a cooler temperature. Because the base pair G+C requires more energy to anneal than the base pair A+T, the rate of reannealing between two strands depends partly on their nucleotide sequence, and it is therefore possible to predict or estimate the sequence of the duplex molecule by the time it takes to fully hybridize. Reassociation kinetics is studied with C0t analysis: fragments reannealing at low C0t values tend to have highly repetitive sequences, while higher C0t values imply more unique sequences.[8]
- receptor
- reciprocal translocation
- A type of chromosomal translocation by which there is a reciprocal exchange of chromosome segments between two or more non-homologous chromosomes. When the exchange of material is evenly balanced, reciprocal translocations are usually harmless.
- recombinant DNA (rDNA)
- Any DNA molecule in which laboratory methods of genetic recombination have brought together genetic material from multiple sources, thereby creating a sequence that would not otherwise be found in a naturally occurring genome. Because DNA molecules from all organisms share the same basic chemical structure and properties, DNA sequences from any species, or even sequences created de novo by artificial gene synthesis, may be incorporated into recombinant DNA molecules. Recombinant DNA technology is widely used in genetic engineering.
- recombinase
- recombination
- See genetic recombination, homologous recombination, and chromosomal crossover.
- recombinator
- Any nucleotide sequence that increases the likelihood of homologous recombination in nearby regions of the genome, e.g. the Chi sequence in certain species of bacteria.[8]
- recon
- The smallest unit of a DNA molecule capable of undergoing homologous recombination, i.e. a pair of consecutive nucleotides, adjacent to each other in cis.[8]
- regulon
- A group of non-contiguous genes which are regulated as a unit, generally by virtue of having their expression controlled by the same regulatory element or set of elements, e.g. the same repressor or activator. The term is most commonly used with prokaryotes, where a regulon may consist of genes from multiple operons.
- repeat
- repetitive DNA
- A region or fragment of DNA consisting largely or entirely of repeated nucleotide sequences.
- replacement mutation
- See nonsynonymous mutation.
- replication
- 1. The process by which certain biological molecules, notably the nucleic acids DNA and RNA, produce copies of themselves.
- 2. A technique used to estimate technical and biological variation in experiments for statistical analysis of microarray data. Replicates may be technical replicates, such as dye swaps or repeated array hybridizations, or biological replicates, biological samples from separate experiments which are used to test the effects of the same experimental treatment.
- replication eye
- The eye-shaped structure that forms when a pair of replication forks, each growing away from the origin, separates the strands of the double helix during DNA replication.
- replication fork
- replication rate
- The speed at which deoxyribonucleotides are incorporated into an elongating chain by DNA polymerases during DNA replication; or more generally the speed at which any chromosome, genome, cell, or organism makes a complete, independently functional copy of itself.
- replicator
- 1. Any fragment or region of DNA that contains a replication origin.[8]
- 2. Any molecule or structure capable of copying itself; namely, nucleic acids, but also crystals of many minerals, e.g. kaolinite.
- replicon
- Any molecule or region of DNA or RNA that replicates from a single origin of replication.
- replisome
- The entire complex of molecular machinery that carries out the process of DNA replication, including all proteins, nucleic acids, and other molecules which participate at an active replication fork.
- reporter
- An MIAME-compliant term to describe a reagent used to make a single measurement in a gene expression experiment. MIAME defines it as "the nucleotide sequence present in a particular location on the array".[5] A reporter may be a segment of single-stranded DNA that is covalently attached to the array surface. Compare probe.
- reporter gene
- repression
- See downregulation.
- repressor
- A DNA-binding protein that inhibits the expression of one or more genes by binding to the operator and blocking the attachment of RNA polymerase to the promoter, thus preventing transcription. This process is known as negative gene regulation.
- rescue
- The restoration of a defective cell or tissue to a healthy or normal condition,[8] or the reversion or recovery of a mutant gene to its normal functionality, especially in the context of experimental genetics, where an experiment (e.g. a drug, cross, or gene transfer) resulting in such a restoration is said to rescue the normal phenotype.
- residue
- An individual monomer or subunit of a larger polymeric macromolecule; e.g. a nucleic acid is composed of nucleotide residues, and a peptide or protein is composed of amino acid residues.[8]
- response element
- A short sequence of DNA within a promoter region that is able to bind specific transcription factors in order to regulate transcription of specific genes.
- restitution
- The spontaneous rejoining of an experimentally broken chromosome which restores the original configuration.
- restitution nucleus
- A nucleus containing twice the expected number of chromosomes owing to an error in cell division, especially an unreduced, diploid product of meiosis resulting from the failure of the first or second meiotic division.
- restriction enzyme
- An endonuclease or exonuclease enzyme that recognizes and cleaves a nucleic acid molecule into fragments at or near specific recognition sequences known as restriction sites by breaking the phosphodiester bonds of the nucleic acid backbone. Restriction enzymes are naturally occurring in many organisms, but are also routinely used for artificial modification of DNA in laboratory techniques such as molecular cloning.
- restriction fragment
- Any DNA fragment that results from the cutting of a DNA strand by a restriction enzyme at one or more restriction sites.
- restriction fragment length polymorphism (RFLP)
- restriction map
- A diagram of known restriction sites within a known DNA sequence, such as a plasmid vector, obtained by systematically exposing the sequence to various restriction enzymes and then comparing the lengths of the resulting fragments, a technique known as restriction mapping. See also gene map.
- restriction site
- A short, specific sequence of nucleotides (typically 4 to 8 bases in length) that is reliably recognized by a particular restriction enzyme. Because restriction enzymes usually bind as homodimers, restriction sites are generally palindromic sequences spanning both strands of a double-stranded DNA molecule. Restriction endonucleases cleave the phosphate backbone between two nucleotides within the recognized sequence itself, but other types of restriction enzymes make their cuts at one end of the sequence or at a nearby sequence.
- reverse genetics
- An experimental approach in molecular genetics in which a researcher starts with a known gene and attempts to determine its function or its effect on phenotype by any of a variety of laboratory techniques, commonly by deliberately mutating the gene's nucleic acid sequence or by repressing or silencing its expression and then screening the mutated organisms for obvious changes in phenotype. When the gene of interest is the only one in the genome whose expression has been manipulated, any observed phenotypic changes are assumed to be influenced by it. This is the opposite of forward genetics, in which a known phenotype is linked to one or more unknown genes.
- reverse transcriptase (RT)
- An enzyme capable of synthesizing a complementary DNA molecule from an RNA template, a process termed reverse transcription.
- reverse transcription
- The synthesis of a DNA molecule from an RNA template, the opposite of ordinary transcription. This process, mediated by the enzyme reverse transcriptase, is used by many viruses to replicate their genomes, as well as by retrotransposons and in eukaryotic cells.
- ribonuclease (RNase)
- Any of a class of nuclease enzymes which catalyze the hydrolytic cleavage of phosphodiester bonds in RNA molecules, thus severing polymeric strands of ribonucleotides into smaller components. Compare deoxyribonuclease.
- ribonucleic acid (RNA)
- A polymeric nucleic acid molecule composed of a series of ribonucleotides which incorporate a set of four nucleobases: adenine (A), guanine (G), cytosine (C), and uracil (U). Unlike DNA, RNA is more often found as a single strand folded onto itself, rather than a paired double strand. Various types of RNA molecules serve in a wide variety of essential biological roles, including coding, decoding, regulating, and expressing genes, as well as functioning as signaling molecules and, in certain viral genomes, as the primary genetic material itself.
- ribonucleotide
- A nucleotide containing ribose as its pentose sugar component, and the monomeric subunit of ribonucleic acid (RNA) molecules. Ribonucleotides canonically incorporate any of four nitrogenous bases: adenine (A), guanine (G), cytosine (C), and uracil (U). Compare deoxyribonucleotide.
- ribonucleotide reductase (RNR)
- An enzyme which catalyzes the formation of deoxyribonucleotides via the reductive dehydroxylation of ribonucleotides, specifically by removing the 2' hydroxyl group from the ribose ring of ribonucleoside diphosphates (rNDPs). RNR plays a critical role in regulating the overall rate of DNA synthesis such that the ratio of DNA to cell mass is kept constant during cell division and DNA repair.
- ribonucleoprotein (RNP)
- A nucleoprotein that is a complex of one or more RNA molecules and one or more proteins. Examples include ribosomes and the enzyme ribonuclease P.
- ribose
- A monosaccharide sugar which, as D-ribose in its pentose ring form, is one of three primary components of the ribonucleotides from which ribonucleic acid (RNA) molecules are built. Ribose differs from its structural analog deoxyribose, used in DNA, only at the 2' carbon, where ribose has an attached hydroxyl group that deoxyribose lacks.
- ribosomal DNA (rDNA)
- ribosomal RNA (rRNA)
- ribosome
- A molecular complex which serves as the site of protein synthesis. Ribosomes consist of two subunits (the small subunit, which reads the messages encoded in mRNA molecules, and the large subunit, which links amino acids in sequence to form a polypeptide chain), each of which is composed of one or more strands of ribosomal RNA and various ribosomal proteins.
- riboswitch
- A regulatory sequence within a messenger RNA transcript that can bind a small effector molecule, preventing or disrupting translation and thereby acting as a switch that regulates the mRNA's expression.
- ribozyme
- An RNA molecule with enzymatic activity, i.e. one that is capable of catalyzing one or more specific biochemical reactions, similar to protein enzymes. Ribozymes function in numerous capacities, including in ribosomes as part of the large subunit ribosomal RNA.
- RNA
- See ribonucleic acid.
- RNA gene
- A gene that codes for any of the various types of non-coding RNA (e.g. rRNA and tRNA).[8]
- RNA interference (RNAi)
- RNA polymerase
- Any of a class of polymerase enzymes that synthesize RNA molecules from a DNA template. RNA polymerases are essential for transcription and are found in all living organisms and many viruses. They build long single-stranded polymers called transcripts by adding ribonucleotides one at a time in the 5'-to-3' direction, relying on the template provided by the complementary strand to transcribe the nucleotide sequence faithfully.
- RNA splicing
- RNA-induced silencing complex (RISC)
- A ribonucleoprotein complex which works to silence endogenous and exogenous genes by participating in various RNA interference pathways at the transcriptional and translational levels. RISC can bind both single-stranded and double-stranded RNA fragments and then cleave them or use them as guides to target complementary mRNAs for degradation.
- RNase
- See ribonuclease.
- Robertsonian translocation (ROB)
- A type of chromosomal translocation by which double-strand breaks at or near the centromeres of two acrocentric chromosomes cause a reciprocal exchange of segments that gives rise to one large metacentric chromosome (composed of the long arms) and one extremely small chromosome (composed of the short arms), the latter of which is often subsequently lost from the cell with little effect because it contains very few genes. The resulting karyotype shows one fewer than the expected total number of chromosomes, because two previously distinct chromosomes have essentially fused together. Carriers of Robertsonian translocations are generally not associated with any phenotypic abnormalities, but do have an increased risk of generating meiotically unbalanced gametes.
- rolling circle replication (RCR)
- rRNA
- See ribosomal RNA.
- rtPCR
- 1. An abbreviation of real-time polymerase chain reaction, synonymous with quantitative PCR.
- 2. An abbreviation of reverse transcription polymerase chain reaction.
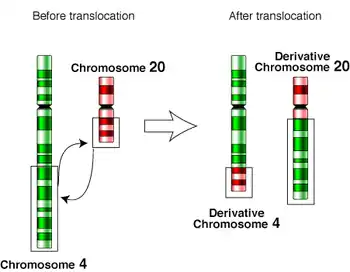
A reciprocal translocation between chromosome 4 and chromosome 20
S
- S phase
- The phase of the cell cycle during which nuclear DNA is replicated, occurring after the G1 phase and before the G2 phase.[4]
- samesense mutation
- See synonymous mutation.
- Sanger sequencing
- A method of DNA sequencing based on the in vitro replication of a DNA template sequence, during which fluorochrome-labeled, chain-terminating dideoxynucleotides are randomly incorporated in the elongating strand; the resulting fragments are then sorted by size with electrophoresis, and the particular fluorochrome terminating each of the size-sorted fragments is detected by laser chromatography, thus revealing the sequence of the original DNA template through the order of the fluorochrome labels as one reads from small-sized fragments to large-sized fragments. Though Sanger sequencing has been replaced in some contexts by next-generation methods, it remains widely used for its ability to produce relatively long sequence reads (500+ nucleotides) and its very low error rate.
- saturation hybridization
- An in vitro nucleic acid hybridization reaction in which one polynucleotide component (either DNA or RNA) is supplied in great excess relative to the other, causing all complementary sequences in the other polynucleotide to pair with the excess sequences and form hybrid duplex molecules.[8]
- scaffolding
- scRNA
- See small conditional RNA.
- selectable marker
- selective sweep
- The process by which strong positive selection of a new and beneficial mutation within a population causes the mutation to reach fixation so quickly that nearby linked DNA sequences also become fixed via genetic hitchhiking, thereby reducing or eliminating the genetic variation of nearby loci within the population.
- selfish genetic element
- Any genetic material (e.g. a gene or any other DNA sequence) which can enhance its own replication and/or transmission into subsequent generations at the expense of other genes in the genome, even if doing so has no positive effect or even a net negative effect on the fitness of the genome as a whole. Selfish elements usually work by producing self-acting gene products which enable them to repeatedly copy and paste themselves into other parts of the genome, independently of normal DNA replication (as with transposable elements); by facilitating the uneven swapping of chromosome segments during genetic recombination events (as with unequal crossing over); or by disrupting the normally equal redistribution of replicated material during mitotis or meiosis such that the probability that the selfish element is present in a given daughter cell is greater than the normal 50 percent (as with gene drives).
- semiconservative replication
- The standard mode of DNA replication that occurs in all living cells, in which each of the two parental strands of the original double-stranded DNA molecule are used as template strands, with DNA polymerases replicating each strand separately and simultaneously in antiparallel directions. The result is that each of the two double-stranded daughter molecules is composed of one of the original parental strands and one newly synthesized complementary strand, such that each daughter molecule conserves the precise sequence of information (indeed the very same atoms) from one-half of the original molecule. Contrast conservative replication and dispersive replication.
- sense
- A distinction made between the individual strands of a double-stranded DNA molecule in order to easily and specifically identify each strand. The two complementary strands are distinguished as sense and antisense or, equivalently, the coding strand and the template strand. It is the antisense/template strand which is actually used as the template for transcription; the sense/coding strand merely resembles the sequence of codons on the RNA transcript, which makes it possible to determine from the DNA sequence alone the expected amino acid sequence of any protein translated from the RNA transcript. Which strand is which is relative only to a particular RNA transcript and not to the entire DNA molecule; that is, either strand can function as the sense/coding or antisense/template strand.
- sense codon
- Any codon that specifies an amino acid, as opposed to a stop codon, which does not specify any particular amino acid but instead signals the end of translation.
- sequence
- See nucleic acid sequence.
- sequence logo
- In bioinformatics, a graphical representation of the conservation of nucleobases or amino acids at each position within a nucleic acid or protein sequence. Sequence logos are created by aligning many sequences and used to depict consensus sequences as well as the degree of variability within the pool of aligned sequences.
- sequence-tagged site (STS)
- Any DNA sequence that occurs exactly once within a particular genome, and whose location and nucleotide sequence are known with confidence.
- sex chromosome
- See allosome.
- sex linkage
- Shine–Dalgarno sequence
- In many prokaryotic messenger RNAs, the consensus sequence AGGAGGU, located 6–8 bases upstream of the translation start codon, which functions as a binding site for the ribosome by complementing a sequence in the ribosomal RNA.[8]
- short arm
- In condensed chromosomes where the positioning of the centromere creates two segments or "arms" of unequal length, the shorter of the two arms of a chromatid. Contrast long arm.
- short tandem repeat (STR)
- See microsatellite.
- short interspersed nuclear element (SINE)
- shotgun sequencing
- silencer
- A sequence or region of DNA that can be bound by a repressor, thereby blocking the transcription of a nearby gene.
- silent allele
- An allele that does not produce a detectable product.[8] Compare null allele.
- silent mutation
- A type of neutral mutation which does not have an observable effect on the organism's phenotype. Though the term "silent mutation" is often used interchangeably with synonymous mutation, synonymous mutations are not always silent, nor vice versa. Missense mutations which result in a different amino acid but one with similar functionality (e.g. leucine instead of isoleucine) are also often classified as silent, since such mutations usually do not significantly affect protein function.
- simple sequence repeat (SSR)
- See microsatellite.
- single-nucleotide polymorphism (SNP)
- Any substitution of a single nucleotide which occurs at a specific position within a genome and with measurable frequency within a population; for example, at a specific base position in a DNA sequence, the majority of the individuals in a population may have a cytosine (C), while in a minority of individuals, the same position may be occupied by an adenine (A). SNPs are usually defined with respect to a "standard" reference genome; an individual human genome differs from the reference human genome at an average of 4 to 5 million positions, most of which consist of SNPs and short indels. See also polymorphism.
- single-strand break (SSB)
- The loss of continuity of the phosphate-sugar backbone in one strand of a DNA duplex.[9] See also nick; contrast double-strand break.
- single-stranded
- Composed of a single, unpaired nucleic acid molecule, i.e. one linear strand of nucleotides sharing a single phosphodiester backbone, as opposed to a duplex of two such strands joined by base pairing. See also single-stranded DNA and single-stranded RNA.
- single-stranded DNA (ssDNA)
- Any DNA molecule that consists of a single nucleotide polymer, or "strand", as opposed to a pair of complementary strands held together by hydrogen bonds (double-stranded DNA). In most circumstances, DNA is more stable and more common in double-stranded form, but high temperatures, low concentrations of dissolved salts, and very high or low pH can cause double-stranded molecules to decompose into two single-stranded molecules in a process known as "melting"; this reaction is exploited by naturally occurring enzymes such as those involved in DNA replication as well as by laboratory techniques such as polymerase chain reaction.
- siRNA
- See small interfering RNA.
- sister chromatids
- A pair of identical copies (chromatids) produced as the result of the DNA replication of a chromosome, particularly when both copies are joined together by a common centromere; the pair of sister chromatids is called a dyad. The two sister chromatids are ultimately separated from each other into two different cells during mitosis or meiosis.
- site-directed mutagenesis
- small conditional RNA (scRNA)
- A class of small RNA molecules engineered so as to change conformation conditionally in response to cognate molecular inputs, often with the goal of controlling signal transduction pathways in vitro or in vivo.
- small interfering RNA (siRNA)
- small nuclear RNA (snRNA)
- small nucleolar RNA (snoRNA)
- small temporal RNA (stRNA)
- A subclass of microRNAs, originally described in nematodes, which regulate the timing of developmental events by binding to complementary sequences in the 3' untranslated regions of messenger RNAs and inhibiting their translation. In contrast to siRNAs, which serve similar purposes, stRNAs bind to their target mRNAs after the initiation of translation and without affecting mRNA stability, which makes it possible for the target mRNAs to resume translation at a later time.
- snoRNA
- See small nucleolar RNA.
- snRNA
- See small nuclear RNA.
- solenoid fiber
- soluble RNA (sRNA)
- See transfer RNA.
- somatic cell
- Any biological cell forming the body of an organism, or, in multicellular organisms, any cell other than a gamete, germ cell, or undifferentiated stem cell. Somatic cells are theoretically distinct from cells of the germ line, meaning the mutations they have undergone can never be transmitted to the organism's descendants, though in practice exceptions do exist.
- somatic cell nuclear transfer (SCNT)
- somatic crossover
- See mitotic recombination.
- Southern blotting
- A molecular biology method used for detecting a specific sequence in DNA samples. The technique combines separation of DNA fragments by gel electrophoresis, transfer of the DNA to a synthetic membrane, and subsequent identification of target fragments with radio-labeled or fluorescent hybridization probes.
- spacer
- Any sequence or region of non-coding DNA separating neighboring genes, whether transcribed or not. The term is used in particular to refer to the non-coding regions between the many repeated copies of the ribosomal RNA genes.[9] See also intergenic region.
- spatially-restricted gene expression
- The expression of one or more genes only within a specific anatomical region or tissue, often in response to a paracrine signal. The boundary between the jurisdictions of two spatially restricted genes may generate a sharp phenotypic gradient there, as with striping patterns.
- spectral karyotype (SKY)
- spindle apparatus
- spliceosome
- splicing
- See genetic engineering.
- split-gene
- ssDNA
- See single-stranded DNA.
- ssRNA
- See single-stranded RNA.
- stable uncharacterized transcript (SUT)
- standard genetic code
- The genetic code used by the vast majority of living organisms for translating nucleic acid sequences into proteins. In this system, of the 64 possible permutations of three-letter codons that can be made from the four nucleotides, 61 code for one of the 20 amino acids, and the remaining three code for stop signals. For example, the codon CAG codes for the amino acid glutamine and the codon UAA is a stop codon. The standard genetic code is described as degenerate or redundant because some amino acids can be coded for by more than one different codon.
- start codon
- The first codon translated by a ribosome from a mature messenger RNA transcript, used as a signal to initiate peptide synthesis. In the standard genetic code, the start codon always codes for the same amino acid, methionine, in eukaryotes and for a modified methionine in prokaryotes. The most common start codon is the triplet AUG. Contrast stop codon.
- statistical genetics
- A branch of genetics concerned with the development of statistical methods for drawing inferences from genetic data. The theories and methodologies of statistical genetics often support research in quantitative genetics, genetic epidemiology, and bioinformatics.
- stem-loop
- stem cell
- Any biological cell which has not yet differentiated into a specialized cell type and which can divide through mitosis to produce more stem cells.
- sticky end
- stop codon
- A codon that signals the termination of protein synthesis during translation of a messenger RNA transcript. In the standard genetic code, three different stop codons are used to dissociate ribosomes from the growing amino acid chain, thereby ending translation: UAG (nicknamed "amber"), UAA ("ochre"), and UGA ("opal"). Contrast start codon.
- strand
- An individual chain of nucleotides comprising a nucleic acid polymer, existing either singly (in which case the nucleic acid molecule is said to be single-stranded) or paired in a duplex (in which case it is said to be double-stranded).
- stringency
- The effect of conditions such as temperature and pH upon the degree of complementarity that is required for a hybridization reaction to occur between two single-stranded nucleic acid molecules. In the most stringent conditions, only exact complements can successfully hybridize; as stringency decreases, an increasing number of mismatches can be tolerated by the two hybridizing strands.[11]
- stRNA
- See small temporal RNA.
- structural gene
- A gene that codes for any protein or RNA product other than a regulatory factor. Structural gene products include enzymes, structural proteins, and certain non-coding RNAs.
- submetacentric
- (of a linear chromosome or chromosome fragment) Having a centromere positioned close to but not exactly in the middle of the chromosome, resulting in chromatid arms of slightly different lengths.[2] Compare metacentric.
- substitution
- A type of point mutation in which a single nucleotide and its attached nucleobase is replaced by a different nucleotide.
- substrate
- 1. A chemical compound or molecule upon which a particular enzyme directly acts, often but not necessarily binding the molecule by forming one or more chemical bonds.[4] See also ligand.
- 2. The substance, biotic or abiotic, upon which an organism grows or lives, or by which it is supported; e.g. a particular growth medium used in cell culture.
- suppression
- See downregulation.
- swivel point
- symporter
- Any of a class of transmembrane transporter proteins which facilitate the transport of two or more different molecules across the membrane at the same time and in the same direction; e.g. glucose and sodium ions. Contrast antiporter and uniporter.
- synapsis
- synaptonemal complex
- syncytium
- syndesis
- The synapsis of chromosomes during meiosis.[8]
- synezis
- The aggregation of chromosomes into a dense knot that adheres to one side of the nucleus, commonly observed during leptonema in certain organisms.[8]
- synonymous mutation
- A type of mutation in which the substitution of one nucleotide base for another results, after transcription and translation, in an amino acid sequence which is identical to the original unmutated sequence. This is possible because of the degeneracy of the genetic code, which allows different codons to code for the same amino acid. Though synonymous mutations are often considered silent, this is not always the case; a synonymous mutation may affect the efficiency or accuracy of transcription, splicing, translation, or any other process by which genes are expressed, and thus become effectively non-silent. Contrast nonsynonymous mutation.
- synteny
- synthesis phase
- See S phase.
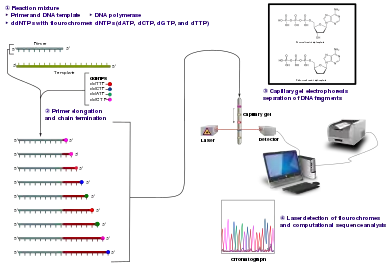
An outline of the Sanger sequencing method
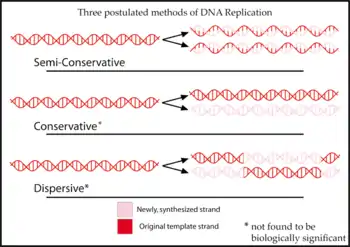
Three different modes of DNA replication. In semiconservative replication, each of the two daughter molecules is built from one of the original parental strands and one newly synthesized strand. In conservative replication, the original parent molecule remains intact while the replicated molecule is composed of two newly synthesized strands. In dispersive replication, each of the daughter molecules is an uneven mix of old and new, with some segments consisting of the two parental strands and others consisting of two newly synthesized strands. Only semiconservative replication occurs naturally.

A sequence logo depicts the statistical frequency with which each nucleobase (or amino acid) occurs within a given sequence. Each position in the sequence is represented by a vertical stack of letters; the total height of the stack indicates the degree of consensus at that position between all of the aligned sequences, and the height of each individual letter in the stack indicates the proportion of the aligned sequences having that nucleobase at that position. A single very large letter filling most of the stack indicates that most or all of the aligned sequences have that particular nucleobase at that position.
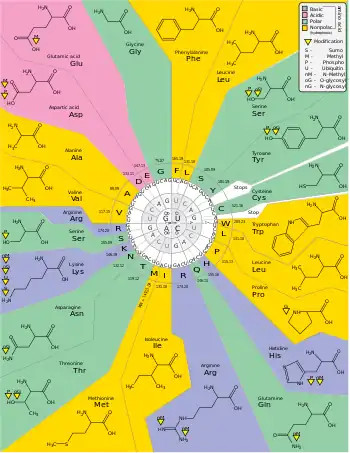
The standard genetic code specifies a set of 20 different amino acids from triplet arrangements of the four different RNA nucleobases (A, G, C, and U). To read this chart, choose one of the four letters in the innermost ring and then move outward, adding two more letters to complete a codon triplet: a total of 64 unique codons can be made this way, 61 of which signal the addition of one of the 20 amino acids (identified by single-letter abbreviation as well as by full name and chemical structure) to a nascent peptide chain, while the remaining three codons are stop codons signalling the termination of translation. Also indicated are some of the chemical properties of the amino acids and the various ways in which they can be modified.
T
- tandem repeat
- A pattern within a nucleic acid sequence in which one or more nucleobases are repeated and the repetitions are directly adjacent (i.e. tandem) to each other. An example is ATGACATGACATGAC, in which the sequence ATGAC is repeated three times.
- TATA box
- A highly conserved non-coding DNA sequence containing a consensus of repeating T and A base pairs that is commonly found in promoter regions of genes in archaea and eukaryotes. The TATA box often serves as the site of initiation of transcription or as a binding site for transcription factors.
- telestability
- Structural destabilization of the DNA double helix at a locus that is relatively distant from the site of binding of a DNA-binding protein.[8]
- telocentric
- (of a linear chromosome or chromosome fragment) Having a centromere positioned at the terminal end of the chromosome (near or within the telomere), resulting in only a single arm.[2] Compare acrocentric.
- telomere
- A region of repetitive nucleotide sequences at each end of a linear chromosome which protects the end of the chromosome from deterioration and from fusion with other chromosomes. Since each round of replication results in the shortening of the chromosome, telomeres act as disposable buffers which are sacrificed to perpetual truncation instead of nearby genes; telomeres can also be lengthened by the enzyme telomerase.
- telomeric silencing
- The repression of transcription of genes in regions adjacent to telomeres. Telomeres also appear to reduce the accessibility of subtelomeric chromatin to modification by DNA methyltransferases.[8]
- telophase
- The final stage of cell division in both mitosis and meiosis, occurring after anaphase and before or simultaneously with cytokinesis, during which a nuclear membrane is synthesized around each set of chromatids, nucleoli are reassembled, and the mitotic spindle is disassembled. Following cytokinesis, the new daughter cells resume interphase.
- template strand
- The strand of a double-stranded DNA molecule which is used as a template for RNA synthesis during transcription. The sequence of the template strand is complementary to the resulting RNA transcript. Contrast coding strand; see also sense.
- terminalization
- In cytology, the progressive shift of chiasmata from their original to more distal positions as meiosis proceeds through diplonema and diakinesis.[8]
- termination codon
- See stop codon.
- terminator
- A DNA sequence or its RNA complement which signals the termination of transcription by triggering processes that ultimately arrest the activity of RNA polymerase or otherwise cause the release of the RNA transcript from the transcriptional complex. Terminator sequences are usually found near the ends of the coding sequences of genes and operons. They generally function after being themselves transcribed into the nascent RNA strand, whereupon the part of the strand containing the sequence either directly interacts with the transcriptional complex or forms a secondary structure such as a hairpin loop which signals the recruitment of enzymes that promote its disassembly.[8]
- three-prime end
- See 3'-end.
- three-prime untranslated region
- See 3' untranslated region.
- thymidine (T, dT)
- One of the four standard nucleosides used in DNA molecules, consisting of a thymine base with its N9 nitrogen bonded to the C1 carbon of a deoxyribose sugar. The prefix deoxy- is commonly omitted, since there are no ribonucleoside analogs of thymidine used in RNA, where it is replaced with uridine instead.
- thymine (T)
- A pyrimidine nucleobase used as one of the four standard nucleobases in DNA molecules. Thymine forms a base pair with adenine. In RNA, thymine is not used at all, and is instead replaced with uracil.
- thymine dimer
- See pyrimidine dimer.
- tissue
- tissue-specific gene expression
- Gene function and expression which is restricted to a particular tissue or cell type. Tissue-specific expression is usually the result of an enhancer which is activated only in the proper cell type.
- topoisomerase
- Any of a class of DNA-binding enzymes which catalyze changes in the topological state of a double-stranded DNA molecule by nicking or cutting the sugar-phosphate backbone of one or both strands, relaxing the torsional stress inherent in the double helix and unwinding or untangling the paired strands before re-ligating the nicks. This process is usually necessary prior to replication and transcription. Topoisomerases thereby convert DNA between its relaxed and supercoiled, linked and unlinked, and knotted and unknotted forms without changing the sequence or overall chemical composition, such that the substrate and product molecules are structural isomers, differing only in their shape and their twisting, linking, and/or writhing numbers.
- totipotency
- trailer sequence
- See 3' untranslated region.
- trait
- trans
- On the opposite side; across from; acting from a different molecule. Contrast cis.
- trans-acting
- Affecting a gene or sequence on a different nucleic acid molecule or strand. A locus or sequence within a particular DNA molecule such as a chromosome is said to be trans-acting if it or its products influence or act upon other sequences located relatively far away or on an entirely different molecule or chromosome. For example, a DNA-binding protein acts "in trans" if it binds to or interacts with a sequence located on any strand or molecule different from the one on which it is encoded. Contrast cis-acting.
- trans-splicing
- transcribed spacer
- A spacer sequence that is transcribed and thus included in the primary ribosomal RNA transcript (as opposed to a non-transcribed spacer) but subsequently excised and discarded during the maturation of functional RNAs of the ribosome.[8]
- transcript
- A product of transcription; that is, any RNA molecule which has been synthesized by RNA polymerase using a complementary DNA molecule as a template. When transcription is completed, transcripts separate from the DNA and become independent primary transcripts. Particularly in eukaryotes, multiple post-transcriptional modifications are usually necessary for raw transcripts to be converted into stable and persistent molecules, which are then described as mature, though not all transcribed RNAs undergo maturation. Many transcripts are accidental, spurious, incomplete, or defective; others are able to perform their functions immediately and without modification, such as certain non-coding RNAs.
- transcript of unknown function (TUF)
- transcriptase
- See RNA polymerase.
- transcription
- The first step in the process of gene expression, in which an RNA molecule, known as a transcript, is synthesized by enzymes called RNA polymerases using a gene or other DNA sequence as a template. Transcription is a critical and fundamental process in all living organisms and is necessary in order to make use of the information encoded within a genome. All classes of RNA must be transcribed before they can exert their effects upon a cell, though only messenger RNA (mRNA) must proceed to translation before a functional protein can be produced, whereas the many types of non-coding RNA fulfill their duties without being translated. Transcription is also not always beneficial for a cell: when it occurs at the wrong time or at a functionless locus, or when mobile elements or infectious pathogens utilize the host's transcription machinery, the resulting transcripts (not to mention the waste of valuable energy and resources) are often harmful to the host cell or genome.
- transcription factor (TF)
- Any protein that controls the rate of transcription of genetic information from DNA to RNA by binding to a specific DNA sequence and promoting or blocking the recruitment of RNA polymerase to nearby genes. Transcription factors can effectively turn "on" and "off" specific genes in order to make sure they are expressed at the right times and in the right places; for this reason, they are a fundamental and ubiquitous mechanism of gene regulation.
- transcription start site (TSS)
- The specific location within a gene at which RNA polymerase begins transcription, defined by the specific nucleotide or codon corresponding to the first ribonucleotide(s) to be assembled in the nascent transcript, i.e. the start codon. This site is usually considered the beginning of the coding sequence and is the reference point for numbering the individual nucleotides within a gene. Nucleotides upstream of the start site are assigned negative numbers and those downstream are assigned positive numbers, which are used to indicate the positions of nearby sequences or structures relative to the TSS. For example, the binding site for RNA polymerase might be a short sequence immediately upstream of the TSS, from approximately -80 to -5, whereas an intron within the coding region might be defined as the sequence starting at nucleotide +207 and ending at nucleotide +793.
- transcription unit
- The segment of DNA between the initiation site and the termination site of transcription, containing the coding sequences for one or more genes. All genes within a transcription unit are transcribed together into a single transcript during a single transcription event; the resulting polycistronic RNA may subsequently be cleaved into separate RNAs, or may be translated as a unit and then cleaved into separate polypeptides.[8]
- transcriptional bursting
- The intermittent nature of transcription and translation mechanisms. Both processes occur in "bursts" or "pulses", with periods of gene activity separated by irregular intervals.
- transcriptome
- The entire set of RNA molecules (often referring to all types of RNA but sometimes exclusively to messenger RNA) that is or can be expressed by a particular genome, cell, population of cells, or species at a particular time or under particular conditions. The transcriptome is distinct from the exome and the translatome.
- transduction
- The transfer of genetic material between cells by a virus or viral vector, either naturally or artificially.
- transfection
- The deliberate experimental introduction of exogenous nucleic acids into a cell or embryo. In the broadest sense the term may refer to any such transfer and is sometimes used interchangeably with transformation, though some applications restrict the usage of transfection to the introduction of naked or purified non-viral DNA or RNA into cultured eukaryotic cells (especially animal cells) resulting in the subsequent incorporation of the foreign DNA into the host genome or the non-hereditary modification of gene expression by the foreign RNA. As a contrast to both standard non-viral transformation and transduction, transfection has also occasionally been used to refer to the uptake of purified viral nucleic acids by bacteria or plant cells without the aid of a viral vector.[8]
- transfer RNA (tRNA)
- A special class of RNA molecule, typically 76 to 90 nucleotides in length, that serves as a physical adapter allowing mRNA transcripts to be translated into sequences of amino acids during protein synthesis. Each tRNA contains a specific anticodon triplet corresponding to an amino acid that is covalently attached to the tRNA's opposite end; as translation proceeds, tRNAs are recruited to the ribosome, where each mRNA codon is paired with a tRNA containing the complementary anticodon. Depending on the organism, cells may employ as many as 41 distinct tRNAs with unique anticodons; because of codon degeneracy within the genetic code, several tRNAs containing different anticodons carry the same amino acid.
- transferase
- transfer-messenger RNA (tmRNA)
- A type of RNA molecule in some bacteria which has dual tRNA-like and mRNA-like properties, allowing it to simultaneously perform a number of different functions during translation.
- transformant
- A cell or organism which has taken up extracellular DNA by transformation and which can express genes encoded by it.
- transformation
- transgene
- Any gene or other segment of genetic material that has been isolated from one organism and then transferred either naturally or by any of a variety of genetic engineering techniques into another organism, especially one of a different species. Transgenes are usually introduced into the second organism's germ line. They are commonly used to study gene function or to confer an advantage not otherwise available in the unaltered organism.
- transition
- A point mutation in which a purine nucleotide is substituted for another purine (A ↔ G) or a pyrimidine nucleotide is substituted for another pyrimidine (C ↔ T). Contrast transversion.
- translation
- The second step in the process of gene expression, in which the messenger RNA transcript produced during transcription is read by a ribosome to produce a functional protein.
- translatome
- The entire set of messenger RNA molecules that are translated by a particular genome, cell, tissue, or species at a particular time or under particular conditions. Like the transcriptome, it is often used as a proxy for quantifying levels of gene expression, though the transcriptome also includes many RNA molecules that are never translated.
- translocation
- A type of chromosomal abnormality caused by the structural rearrangement of large sections of one or more chromosomes. There are two main types: reciprocal and Robertsonian.
- transmembrane protein
- transmission genetics
- The branch of genetics that studies the mechanisms involved in the transfer of genes from parents to offspring.[8]
- transposable element (TE)
- transposase
- Any of a class of self-acting enzymes capable of binding to the flanking sequences of the transposable element which encodes them and catalyzing its movement to another part of a genome, typically by an excision/insertion mechanism or a replicative mechanism, in a process known as transposition.
- transposition
- The process by which a nucleic acid sequence known as a transposable element changes its position within a genome, either by excising and re-inserting itself at a different locus (cut-and-paste) or by duplicating itself without moving the element from its original locus (copy-paste). These reactions are catalyzed by an enzyme known as a transposase which is encoded by a gene within the transposable element itself; thus the element's products are self-acting and can autonomously direct their own reproduction. Transposed sequences may re-insert at random loci or at sequence-specific targets, either on the same DNA molecule or on different molecules.
- transvection
- transversion
- A point mutation in which a purine nucleotide is substituted for a pyrimidine nucleotide, or vice versa (e.g. A ↔ C or A ↔ T). Contrast transition.
- tricarboxylic acid cycle (TCA)
- See citric acid cycle.
- trinucleotide repeat
- Any sequence in which an individual nucleotide triplet is repeated many times in tandem, whether in a gene or non-coding sequence. At most loci some degree of repetition is normal and harmless, but mutations which cause specific triplets (especially those of the form CnG) to increase in copy number above the normal range are highly unstable and responsible for a variety of genetic disorders.
- triplet
- A unit of three successive nucleotides in a DNA or RNA molecule.[8] A triplet within a coding sequence that codes for a specific amino acid is known as a codon.
- trisomy
- A type of polysomy in which a diploid cell or organism has three copies of a particular chromosome instead of the normal two.
- tRNA
- See transfer RNA.
- tRNA-ligase
- See aminoacyl-tRNA synthetase.
- twisting number

A simplified diagram of transcription. RNA polymerase (RNAP) synthesizes an RNA transcript (blue) in the 5'-to-3' direction, using one of the DNA strands as a template, while a complex of multiple transcription factors binds to a promoter upstream of the gene.
U
- ubiquitination
- umber
- See opal.
- uncharged tRNA
- A transfer RNA without an attached amino acid. Contrast charged tRNA.
- underdominance
- underwinding
- See negative supercoiling.
- unequal crossing over
- uniparental inheritance
- unique DNA
- A class of DNA sequences determined by C0t analysis to be present only once in the analyzed genome, as opposed to repetitive sequences. Most structural genes and their introns are unique.[8]
- unstable mutation
- A mutation with a high frequency of reversion.[8]
- untranslated region (UTR)
- Either of two non-coding sequences which are transcribed along with a protein-coding sequence, and thus included within a messenger RNA, but which are not ultimately translated during protein synthesis. A typical mRNA transcript includes one such region immediately upstream of the coding sequence, known as the 5' untranslated region (5'-UTR), and one downstream of the coding sequence, known as the 3' untranslated region (3'-UTR). These regions are not removed during post-transcriptional processing (unlike introns) and are usually considered exclusive of the 5' cap and the 3' polyadenylated tail (both of which are later additions to a primary transcript and not themselves products of transcription). UTRs are a consequence of the fact that transcription usually begins considerably upstream of the start codon of the protein-coding sequence and terminates long after the stop codon has been transcribed, whereas translation is more precise.
- upregulation
- Any process, natural or artificial, which increases the level of gene expression of a certain gene. A gene which is observed to be expressed at relatively high levels (such as by detecting higher levels of its mRNA transcripts) in one sample compared to another sample is said to be upregulated. Contrast downregulation.
- upstream
- Towards or closer to the 5'-end of a chain of nucleotides, or the N-terminus of a peptide chain. Contrast downstream.
- upstream activating sequence (UAS)
- uracil (U)
- A pyrimidine nucleobase used as one of the four standard nucleobases in RNA molecules. Uracil forms a base pair with adenine. In DNA, uracil is not used at all, and is instead replaced with thymine.
- uridine (U, Urd)
- One of the four standard nucleosides used in RNA molecules, consisting of a uracil base with its N9 nitrogen bonded to the C1 carbon of a ribose sugar. In DNA, uridine is replaced with thymidine.
V
- vacuole
- variable number tandem repeat (VNTR)
- variegation
- Variation or irregularity in a particular phenotype, especially a conspicuous visible trait such as color or pigmentation, occurring simultaneously in different parts of the same individual organism due to any of a variety of causes, such as X-inactivation, mitotic recombination, transposable element activity, position effects, or infection by pathogens.
- variome
- vector
- Any DNA molecule used as a vehicle to artificially transport foreign genetic material into another cell, where it can be replicated and/or expressed. Vectors are typically engineered recombinant DNA sequences consisting of an insert (often a transgene) and a longer "backbone" sequence containing an origin of replication, a multiple cloning site, and a selectable marker. Vectors are widely used in molecular biology laboratories to isolate, clone, or express the insert in the target cell.
- vegetal cell
- See somatic cell.
- vesicle
W
- Warburg effect
- western blotting
- whole genome sequencing (WGS)
- The process of determining the entirety or near-entirety of the DNA sequences comprising an organism's genome with a single procedure or experiment, generally inclusive of all chromosomal and extrachromosomal (e.g. mitochondrial) DNA.
- wild type (WT)
- The phenotype of the typical form of a species as it occurs in nature; a product of the standard "normal" allele at a given locus, as opposed to that produced by a non-standard mutant allele.
- wobble base pairing
- writhing number
X
- X chromosome
- One of two sex chromosomes present in organisms which use the XY sex-determination system, and the only sex chromosome in the X0 system. The X chromosome is found in both males and females and typically contains much more gene content than its counterpart, the Y chromosome.
- X-inactivation
- The process by which one of the two copies of the X chromosome is silenced by being irreversibly condensed into transcriptionally inactive heterochromatin in the cells of female therian mammals. A form of dosage compensation, X-inactivation prevents females from producing twice as many gene products from genes on the X chromosome as males, who only have one copy of the X chromosome. Which X chromosome is inactivated is randomly determined in the early embryo, making it possible for cell lineages with different inactive Xs to exist in the same organism.
- X-linked trait
Y
- Y chromosome
- One of two sex chromosomes present in organisms which use the XY sex-determination system. The Y chromosome is found only in males and is typically much smaller than its counterpart, the X chromosome.
- Y fork
- See replication fork.
- yeast artificial chromosome (YAC)
Z
- Z-DNA
- zinc finger
- zygonema
- In meiosis, the second of five substages of prophase I, following leptonema and preceding pachynema. During zygonema, synapsis occurs, physically binding homologous chromosomes to each other, and the cell's centrosome divides into two daughter centrosomes, each containing a single centriole.[8]
- zygosity
- The degree to which multiple copies of a gene, chromosome, or genome have the same genetic sequence; e.g. in a diploid organism with two complete copies of its genome (one maternal and one paternal), the degree of similarity of the alleles present in each copy. Individuals carrying two different alleles for a particular gene are said to be heterozygous for that gene; individuals carrying two identical alleles are said to be homozygous for that gene. Zygosity may also be considered collectively for a group of genes, or for the entire set of genes and genetic loci comprising the genome.
- zygote
- A type of eukaryotic cell formed as the direct result of a fertilization event between two gametes. In multicellular organisms, the zygote is the earliest developmental stage.
See also
References
- "Talking Glossary of Genetic Terms". genome.gov. 8 October 2017. Retrieved 8 October 2017.
- Klug, William S.; Cummings, Michael R. (1986). Concepts of Genetics (2nd ed.). Glenview, Ill.: Scott, Foresman and Company. ISBN 0-673-18680-6.
- Rayner TF; Rocca-Serra P; Spellman PT; Causton HC; et al. (2006). "A simple spreadsheet-based, MIAME-supportive format for microarray data: MAGE-TAB". BMC Bioinformatics. 7: 489. doi:10.1186/1471-2105-7-489. PMC 1687205. PMID 17087822.
- MacLean, Norman (1987). Dictionary of Genetics & Cell Biology. New York: New York University Press. ISBN 0-8147-5438-4.
- Oliver S (2003). "On the MIAME Standards and Central Repositories of Microarray Data". Comparative and Functional Genomics. 4 (1): 1. doi:10.1002/cfg.238. PMC 2447402. PMID 18629115.
- Brazma A (2009). "Minimum Information About a Microarray Experiment (MIAME)--successes, failures, challenges". ScientificWorldJournal. 9: 420–3. doi:10.1100/tsw.2009.57. PMC 5823224. PMID 19484163.
- Functional Genomics Data Society (June 2012). "Minimum Information about a high-throughput SEQuencing Experiment".
- King, Robert C.; Stansfield, William D.; Mulligan, Pamela K. (2006). A Dictionary of Genetics (7th ed.). Oxford: Oxford University Press. ISBN 978-0-19-530762-7.
- Rieger, Rigomar (1991). Glossary of Genetics: Classical and Molecular (5th ed.). Berlin: Springer-Verlag. ISBN 3540520546.
- "7.6D: The Incorporation of Nonstandard Amino Acids". LibreTexts Biology.
- Lewin, Benjamin (2003). Genes VIII. Upper Saddle River, NJ: Pearson Prentice Hall. ISBN 0-13-143981-2.
- "overexpression". Oxford Living Dictionary. Oxford University Press. 2017. Archived from the original on February 10, 2018. Retrieved 18 May 2017.
The production of abnormally large amounts of a substance which is coded for by a particular gene or group of genes; the appearance in the phenotype to an abnormally high degree of a character or effect attributed to a particular gene.
- "overexpress". NCI Dictionary of Cancer Terms. National Cancer Institute at the National Institutes of Health. 2011-02-02. Retrieved 18 May 2017.
overexpress
In biology, to make too many copies of a protein or other substance. Overexpression of certain proteins or other substances may play a role in cancer development.
Further reading
- Budd, A. (2012). "Introduction to genome biology: features, processes, and structures". Evolutionary Genomics. Methods in Molecular Biology. Vol. 855. pp. 3–4. doi:10.1007/978-1-61779-582-4_1. ISBN 978-1-61779-581-7. PMID 22407704.
This article is issued from Wikipedia. The text is licensed under Creative Commons - Attribution - Sharealike. Additional terms may apply for the media files.